
Fine and ultrafine particle toxicity
ExposomeMap-PM

Online access and exploration: https://markopolo.elixir-luxembourg.org
Development status: First version is complete and available for exploration online
Diseases: Multiple related diseases
Sustainable support: Luxembourg Institute of Health, MINERVA
Construction tools: yEd Graph Editor, CellDesigner
Funding: Grant Agreement No. 101156161, https://markersofpollution-markopolo.eu/
License: Creative Commons Attribution 4.0 International (CC BY 4.0) License
Contact: Petr Nazarov, Luxembourg Institute of Health,
Open in MINERVA
Guide to overlays
Colour-coded
overview
Description
The ExposomeMap-PM effort is initiated within the MARKOPOLO project that aims to identify disease-relevant biomarkers and causality mechanisms to understand the biological pathways of cerebral, pulmonary and cardiovascular health outcomes towards improving risk assessment and allowing evaluation of the effectiveness of mitigation strategies. The ExposomeMap-PM is an open, FAIR-compliant model that connects particle exposure to biological mechanisms and health outcomes. The model highlights critical steps such as oxidative stress, inflammation and barrier disruption, which together contribute to various conditions including asthma, chronic obstructive pulmonary disease, cardiovascular and neurodegenerative diseases.
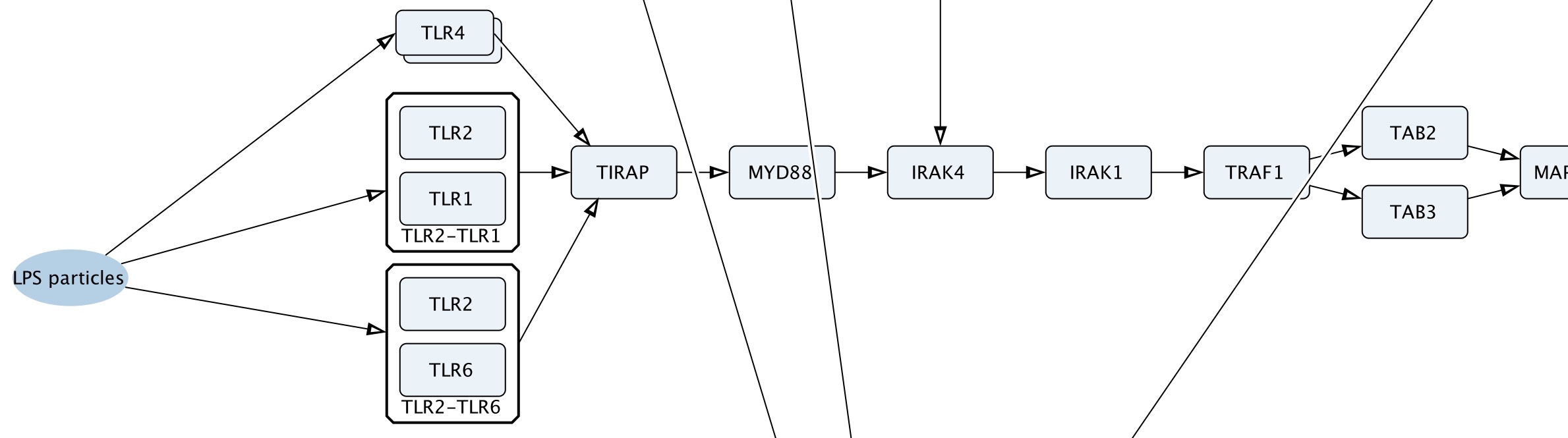
Figure 1. A fragment of the ExposomeMap-UFP showing direct interaction of LPS-containing particles with Toll-like receptors.
Related conditions include respiratory diseases: asthma (DOID:2841), COPD (DOID:3083); cardiovascular diseases: stroke (DOID:6713), heart attack (DOID:5844), hypertension (DOID:10763); neurodegenerative diseases: Alzheimer’s disease (DOID:10652), Parkinson’s disease (DOID:14330), dementia (DOID:1307); skin diseases: atopic dermatitis (DOID:2723); degenerative joint diseases: osteoarthritis (DOID:8398); autoimmune diseases: rheumatoid arthritis (DOID:7148), multiple sclerosis (DOID:2377), inflammatory bowel disease (DOID:0050589).
Omics data visualisation
A step-by-step guide is available to show how an overlay can be created by uploading omics dataset to the map.
Blocks of the map (colour-coded overview)
The map is organised into five blocks shown in the colour-coded overview: receptor activation (purple), receptor-mediated uptake (blue), ion channel perturbation (red), oxidative and catalytic interactions (green) and antioxidant response (beige). They represent groups of the early molecular events triggered by particle deposition on epithelial and immune cells.
Supporting references
ExposomeMap-PM v1.0 was developed based on a structured review of the literature on particulate matter exposure and its molecular, cellular and systemic effects. The full list of references used to construct the map is available in the reference collection.
Editorial Panel
 |
Andreas Daiber, PhD University Medical Center of the Johannes Gutenberg University Mainz, Germany Univ.-Prof., Head of Laboratory of Molecular Cardiology, Scientific Coordinator of MARKOPOLO |
 |
Marin Kuntic, PhD University Medical Center of the Johannes Gutenberg University Mainz, Germany Postdoctoral Researcher |
 |
Thomas Münzel, MD University Medical Center of the Johannes Gutenberg University Mainz, Germany Head of the Cardiology Department |
 |
Jos Lelieveld, PhD Max Planck Institute for Chemistry, Mainz, Germany Director of the Atmospheric Chemistry Department Climate and Atmosphere Research Center, The Cyprus Institute, Nicosia, Cyprus Professor |
 |
Katrin Frauenknecht, MD University Medical Center of the Johannes Gutenberg University Mainz, Germany Consultant Neuropathologist/Deputy Director |
 |
Katja Kanninen, PhD A.I. Virtanen Institute for Molecular Sciences, University of Eastern Finland, Finland Professor of Cellular Neurobiology |
 |
Thuy Thi Lai, PhD A.I. Virtanen Institute for Molecular Sciences, University of Eastern Finland, Finland Postdoctoral Researcher |
 |
Adelina Rogowska-Wrzesinska, PhD University of Southern Denmark, Denmark Associate Professor at the Department of Biochemistry and Molecular Biology |
 |
Amalie Schufri Klinkby, MSc University of Southern Denmark, Denmark PhD Student |
 |
Bharath Anila Bhuvanendran Nair, PhD University of Southern Denmark, Denmark Postdoc |
Development Team
 |
Alexander Mazein, PhD Luxembourg Institute of Health, Luxembourg Postdoctoral Fellow at the Multi-omics Data Science group |
 |
Corrado Ameli, PhD Luxembourg Institute of Health, Luxembourg Postdoctoral Fellow at the Multiomics Data Science group |
 |
Salla Akerblom, BSc Luxembourg Institute of Health, Luxembourg; University of Lille, France MSc Student in Bioinformatics |
 |
Claudia Gutierrez Ortiz, MSc Luxembourg Institute of Health, Luxembourg PhD Student at Socio-Economic & Environmental Health & Health Services |
 |
Maryna Chepeleva, MSc Luxembourg Institute of Health, Luxembourg PhD Student at the Multiomics Data Science group |
 |
Benoit Kunath, PhD Luxembourg Institute of Health, Luxembourg Postdoctoral Fellow at the Bioinformatics & AI group |
 |
Lu Zhang, MSc Luxembourg Institute of Health, Luxembourg Research Engineer at the Bioinformatics & AI group |
 |
Marek Ostaszewski, PhD University of Luxembourg, Luxembourg Research Scientist at the Luxembourg Centre for Systems Biomedicine |
 |
Ahmed Hemedan, PhD Luxembourg Institute of Health, Luxembourg Postdoctoral Fellow at the Department of Transversal Translational Medicine |
 |
Vladimir Despotovic, PhD Luxembourg Institute of Health, Luxembourg Scientist in Health Data, Bioinformatics & AI |
 |
Reka Toth, PhD Luxembourg Institute of Health, Luxembourg Group Leader, Multiomics Data Science; Scientist, Bioinformatics & AI |
 |
Petr Nazarov, PhD Luxembourg Institute of Health, Luxembourg Head of the Bioinformatics and AI Unit |
Acknowledgements
The project is developing in collaboration with the Luxembourg Centre for Systems Biomedicine.

|
This project is supported by ELIXIR Luxembourg (ELIXIR-LU) Node. ELIXIR-LU hosts and maintains the MINERVA Platform for this project and supports its development. |
Funding
This work is supported by the environmental research consortium MARKOPOLO under the HORIZON call HLTH-2024-ENVHLTH-02-06 (Grant Agreement Number 101156161) funded by the European Union and the Swiss State Secretariat for Education, Research and Innovation (SERI).