
Map of ulcerative colitis and atopic dermatitis
Online access and exploration: https://imi-immuniverse.elixir-luxembourg.org
Development status: Available for exploration online
Diseases: Ulcerative colitis and atopic dermatitis
Disease IDs | Ulcerative colitis: DOID:8577, MESH:D003093, MONDO:0005101
Disease IDs | Atopic dermatitis: DOID:3310, MESH:D003876, MONDO:0011292
Sustainable support: LCSB, MINERVA Platform
Construction tool: CellDesigner
License: Creative Commons Attribution 4.0 International (CC BY 4.0) License
Funding: IMI2 ImmUniverse No 853995, ImmUniverse
Contact: Oxana Lopata, University of Luxembourg, Luxembourg, oxana.lopata@uni.lu
Description
The IMIDs Map is accessible here. It was developed within the IMI2 ImmUniverse project. The ImmUniverse project focuses on ulcerative colitis and atopic dermatitis towards new insights into disease severity and progression with the ultimate aim of improved diagnostic and therapeutic options for patients.
How to cite
Lopata O, Acencio ML, Wang X, Hemedan AA, Chao MJ, Jelinsky SA, Tran F, Rosenstiel P, Li Yim AYF, Schneider R, Satagopam V, Ostaszewski M. Identification of shared and unique mechanisms of atopic dermatitis and ulcerative colitis by construction and computational analysis of disease maps. Comput Struct Biotechnol J. 2025 Sep 7;27:4007-4018. doi: 10.1016/j.csbj.2025.09.008. PMID: 41017825.
Funding
This project has received funding from the Innovative Medicines Initiative 2 Joint Undertaking (JU) under grant agreement No 853995.
Acknowledgements

|
This project is supported by ELIXIR Luxembourg (ELIXIR-LU) Node. ELIXIR-LU hosts and maintains the MINERVA Platform for this project and supports its development. |
User guide
This quick user guide explains how to navigate and use the UC and AD disease maps.
The main entry point to the maps is https://imi-immuniverse.elixir-luxembourg.org. This is a comparison diagram of Ulcerative Colitis (UC) and Atopic Dermatitis (AD) disease mechanisms.
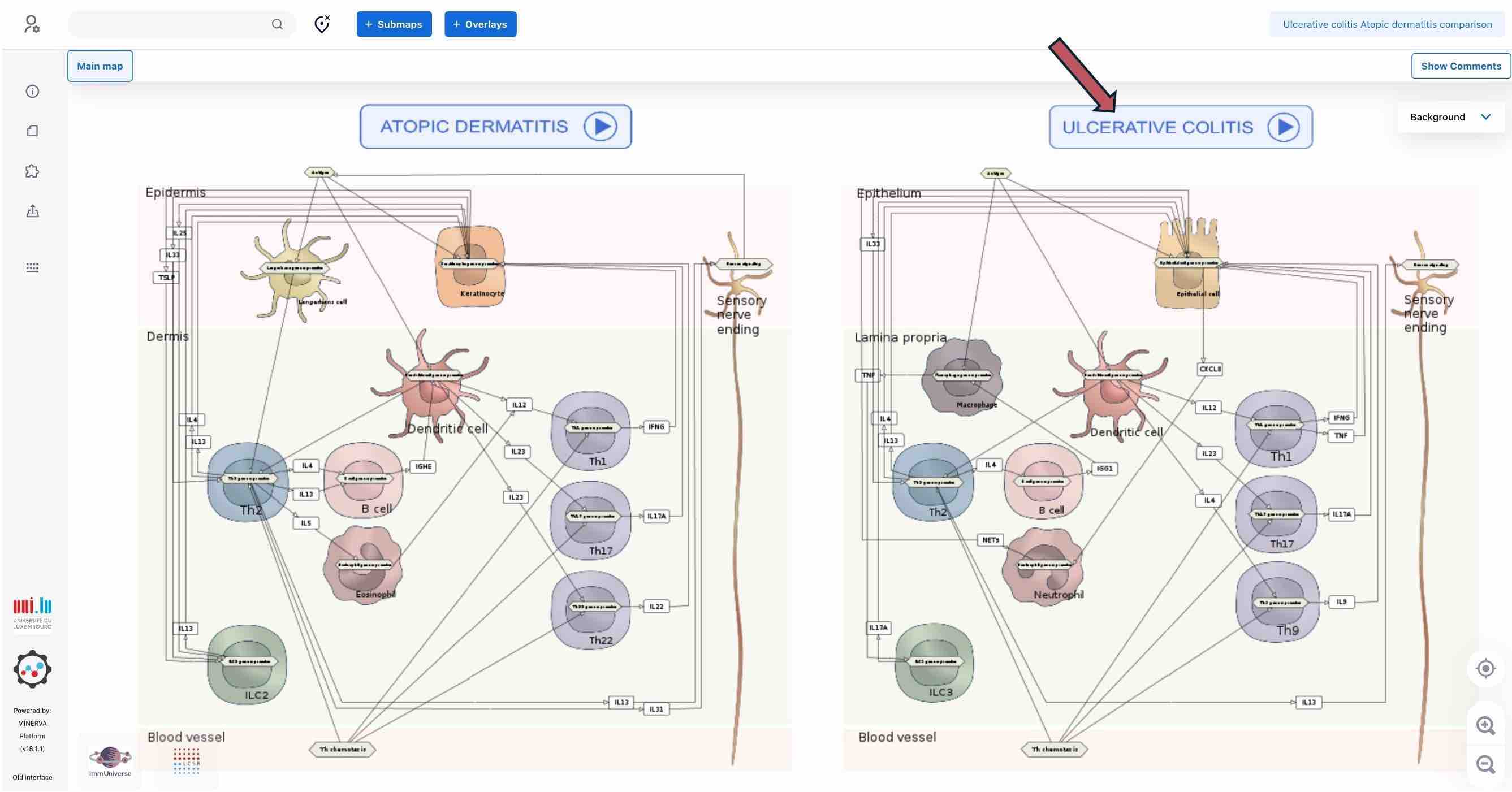
To access detailed maps for both diseases, please click on the blue button with disease name (red arrow) above each part of the diagram.
For example, clicking on the ULCERATIVE COLITIS button you will be directed to the UC intercellular map.
Pan and zoom
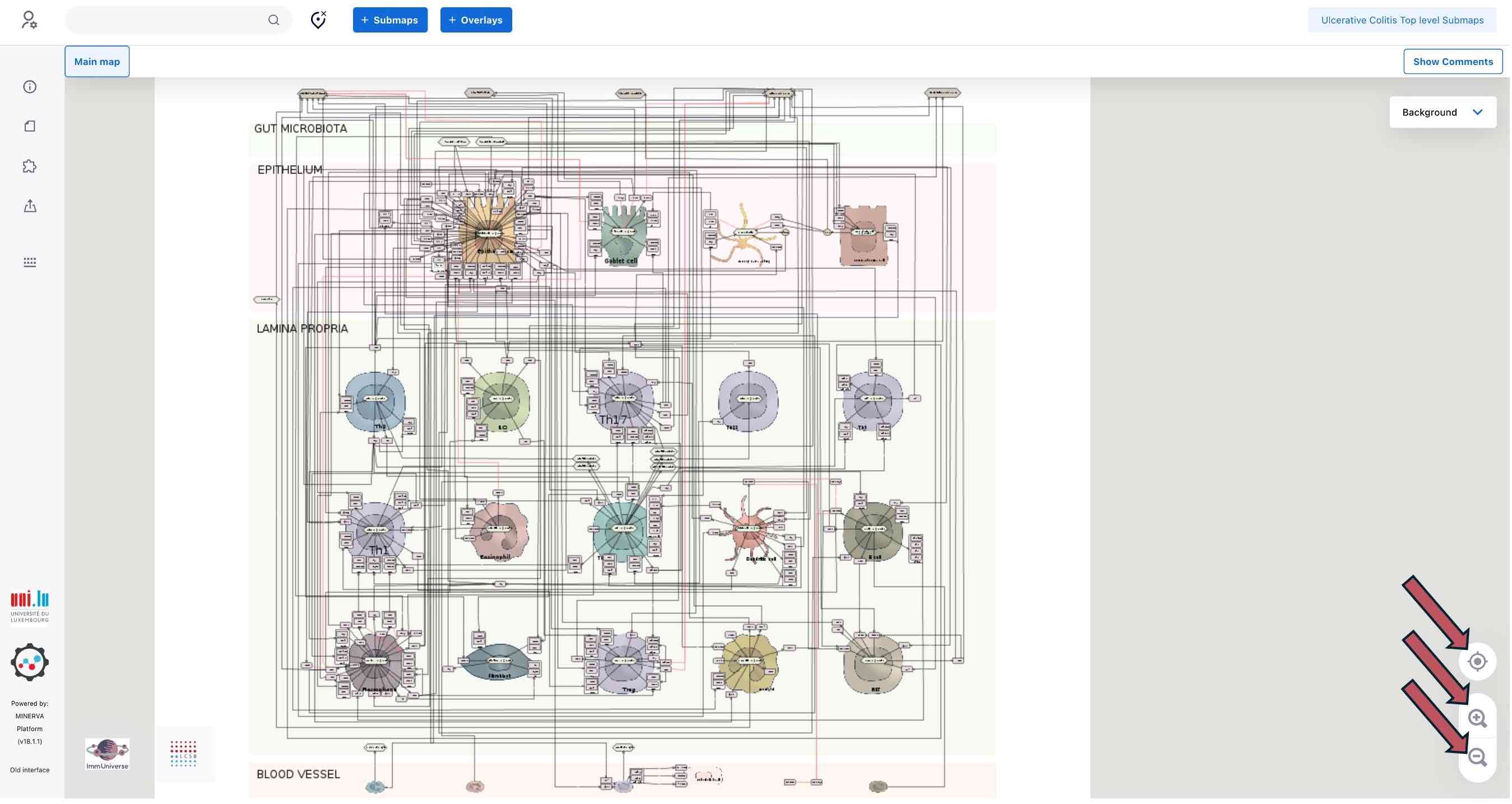
There, you can browse, search in the content or explore data overlays. Zooming and navigating for browsing the map content are similar to working with Google Maps. You can use buttons on the right (red arrows) to zoom in, zoom out and center the map content.
Interactive exploration
To access information about map elements - cell types, phenotypes, proteins, genes, interactions - please click on the element and check the appearing panel with the description on the left.
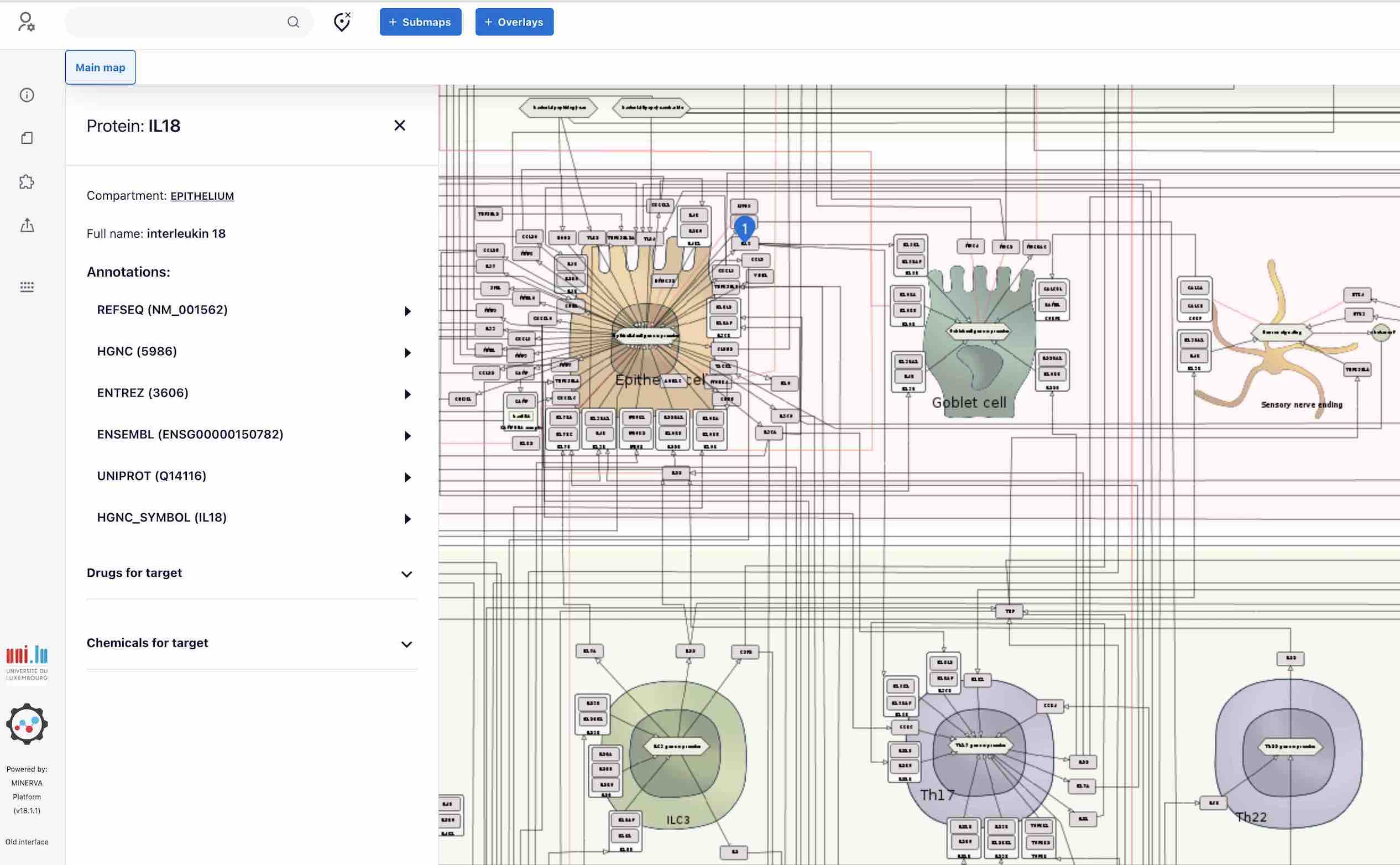
By clicking on a map interaction (black or red lines that connect map elements) you can access references to publications supporting this reaction.
Search
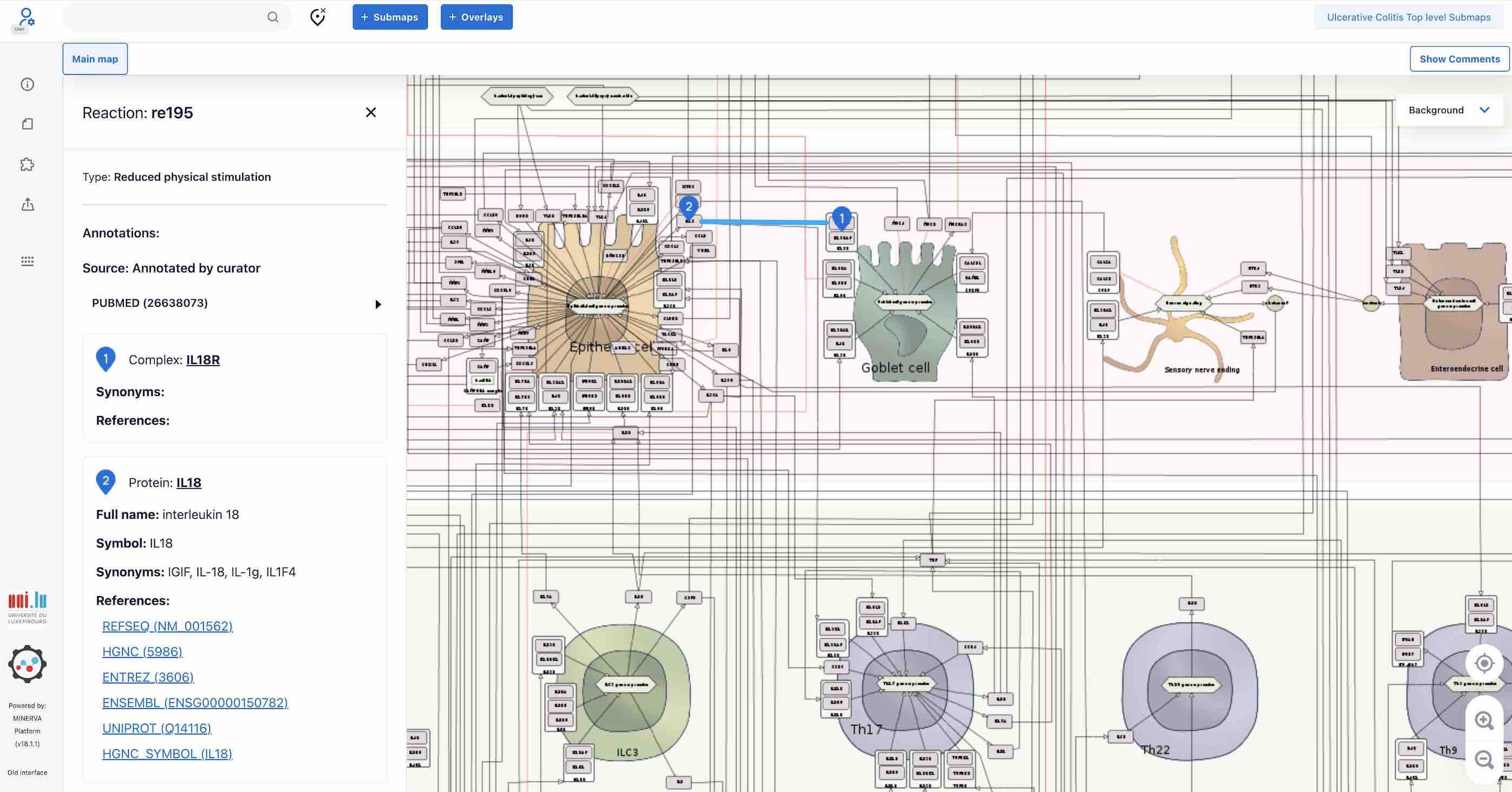
You can search for elements using the search panel and see them in the map highlighted with blue anchors. To clear the search, please use the Clear button
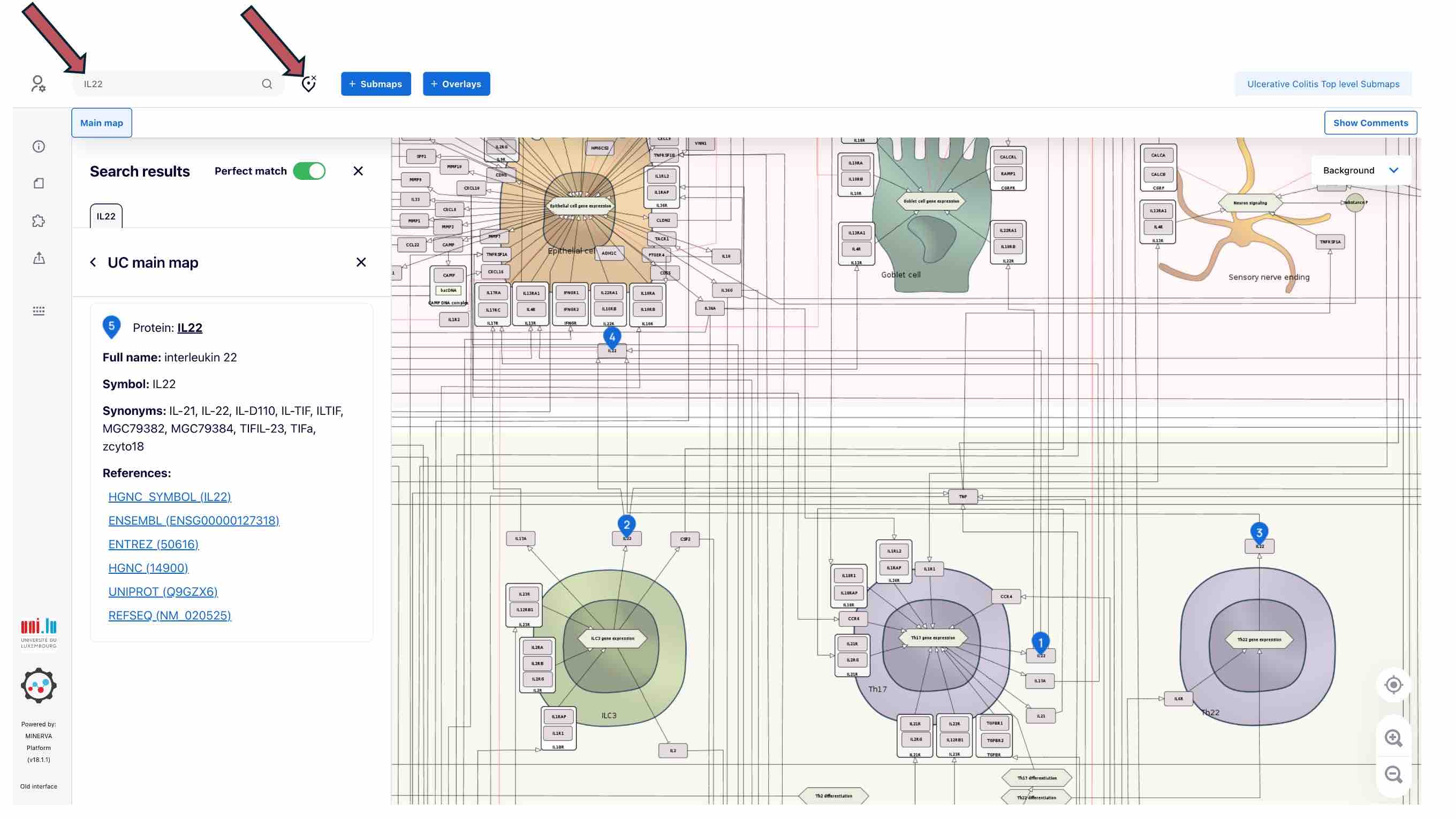
Submaps
To go to the next layer - submaps of UC or AD key cell types - click the SUBMAPS button above. You will see the list of the submaps on the left panel.
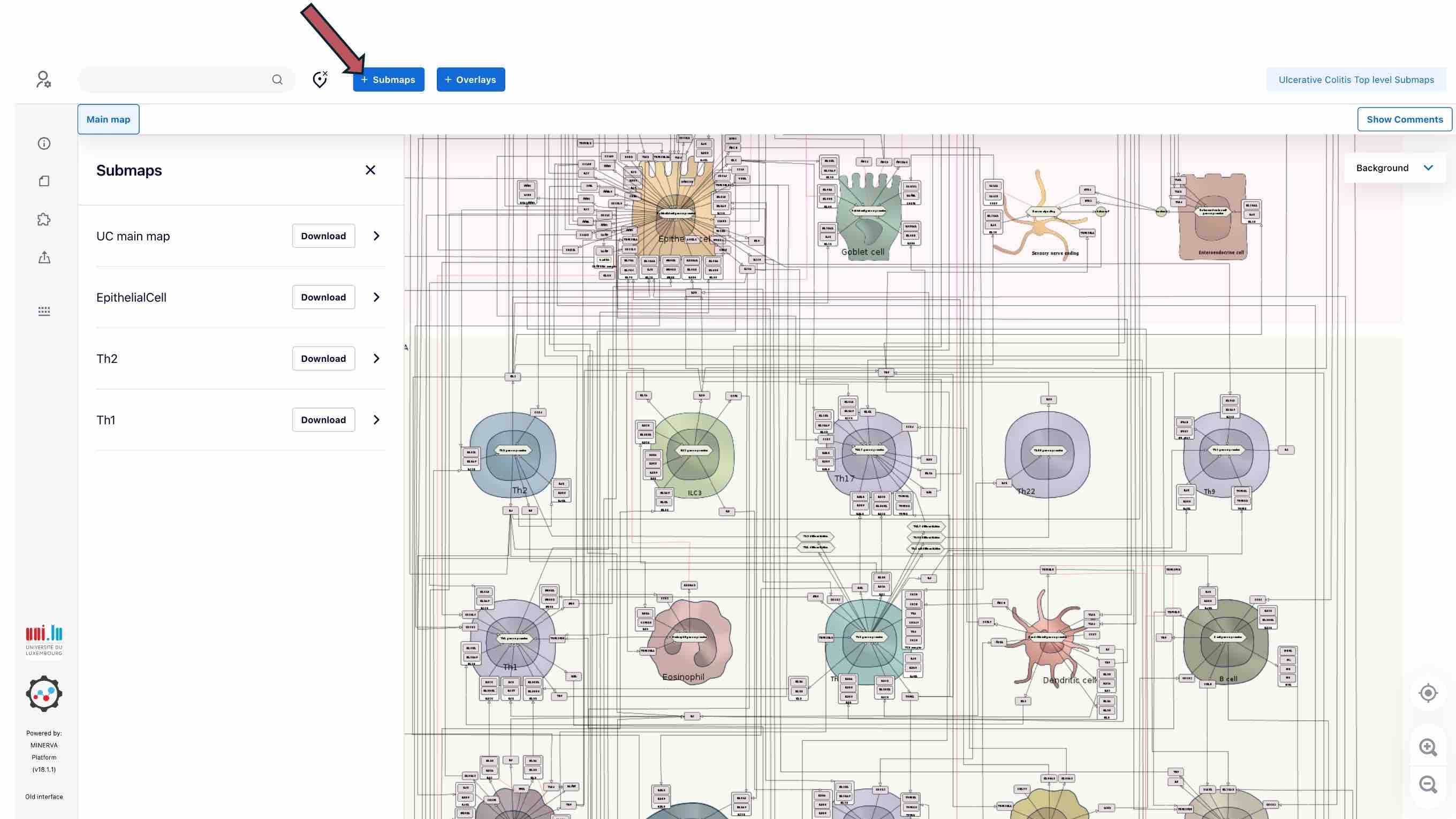
By clicking on the submap name you will be accessing the corresponding submap.
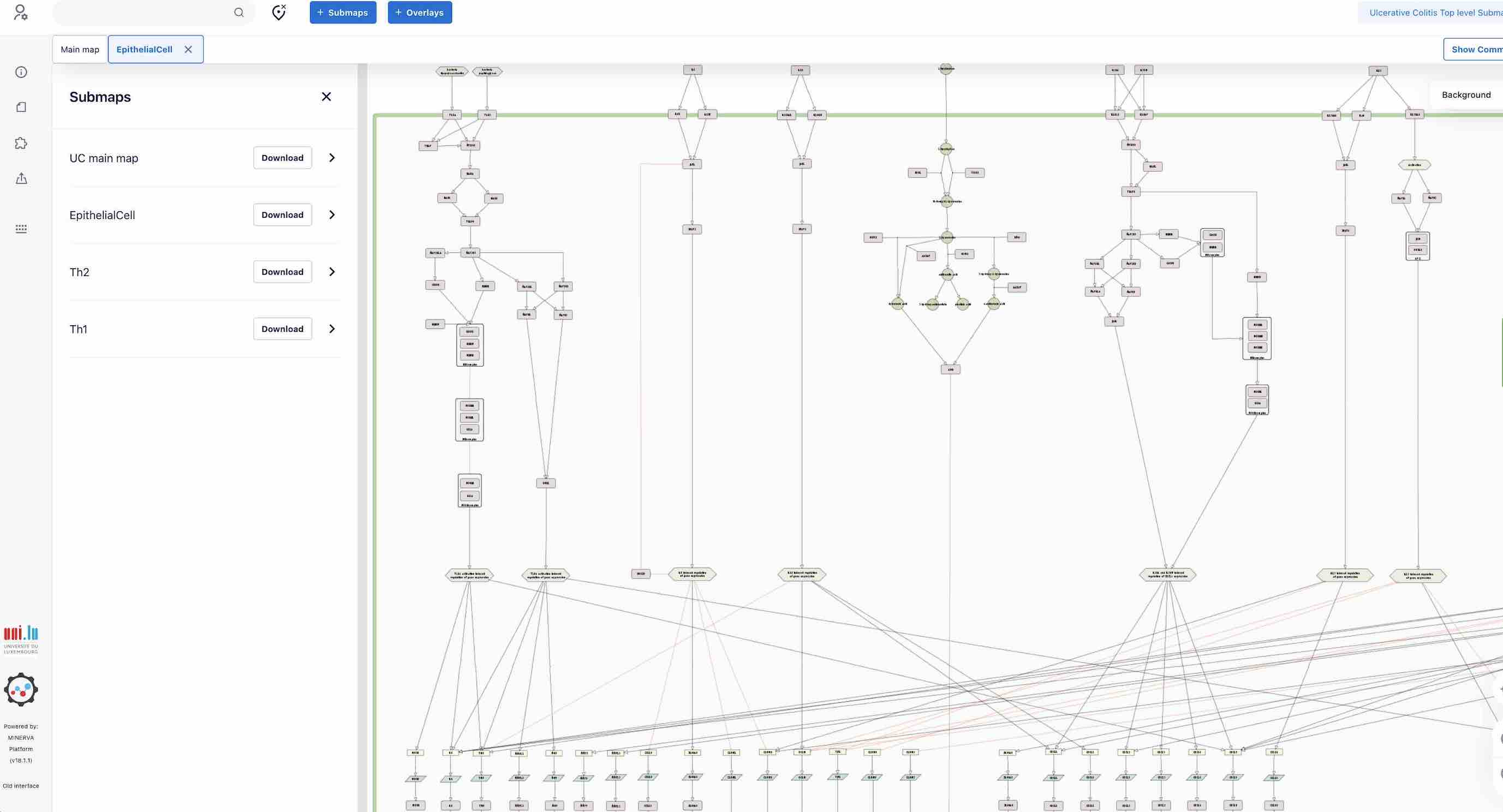 Here you can also search for elements, check description and references etc.
Here you can also search for elements, check description and references etc.
Overlays
To visualise experimental dataset on the map, please use the OVERLAYS button above.
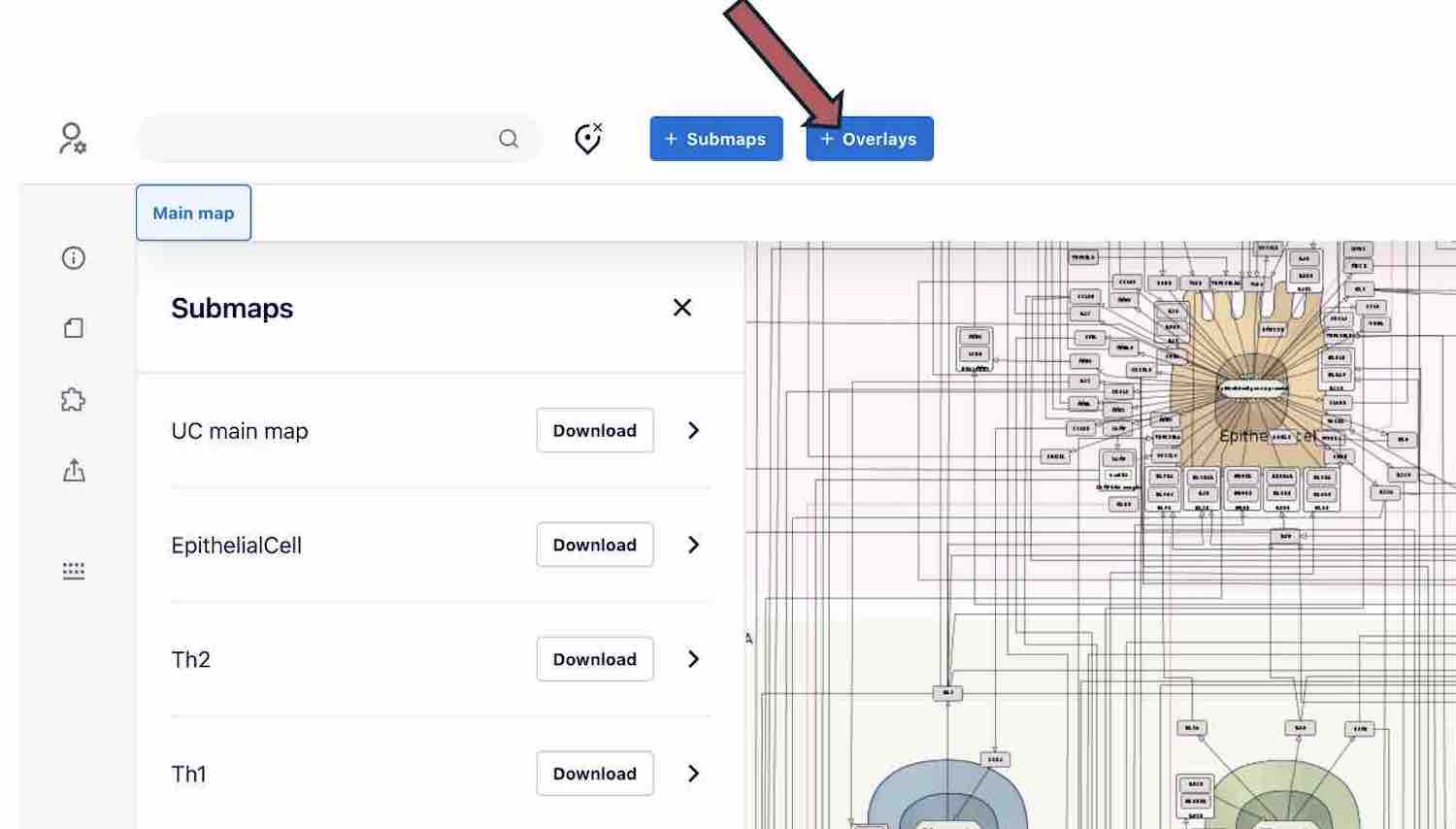
For example, here are genes associated with UC and AD from the Open Targets Genetics database and ExpressionAtlas formatted and highlighted with blue and red to visualize overlaps with disease mechanisms represented in the maps.
Use buttons View and Hide to display the dataset. Many datasets can be shown at the same time.
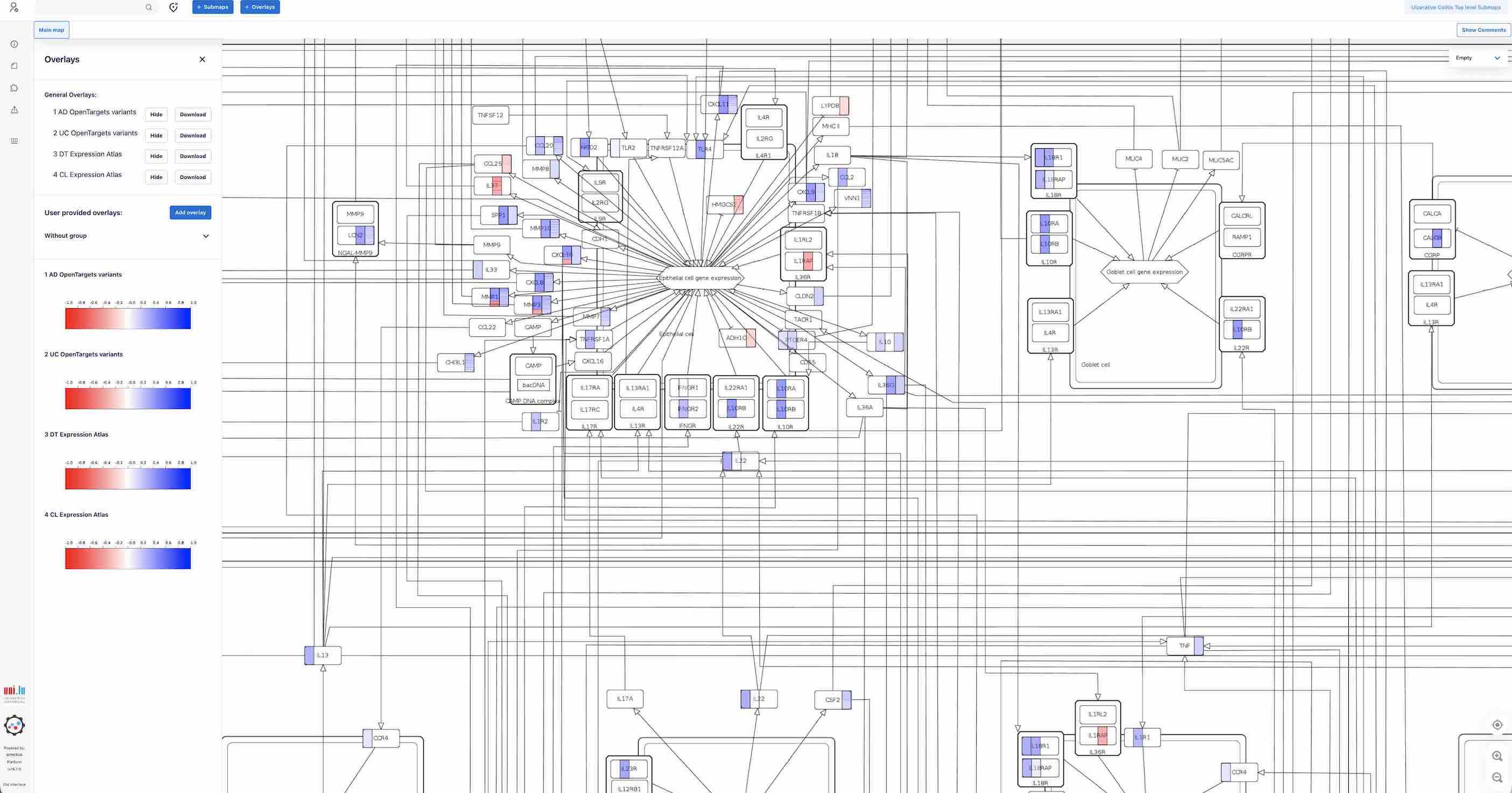
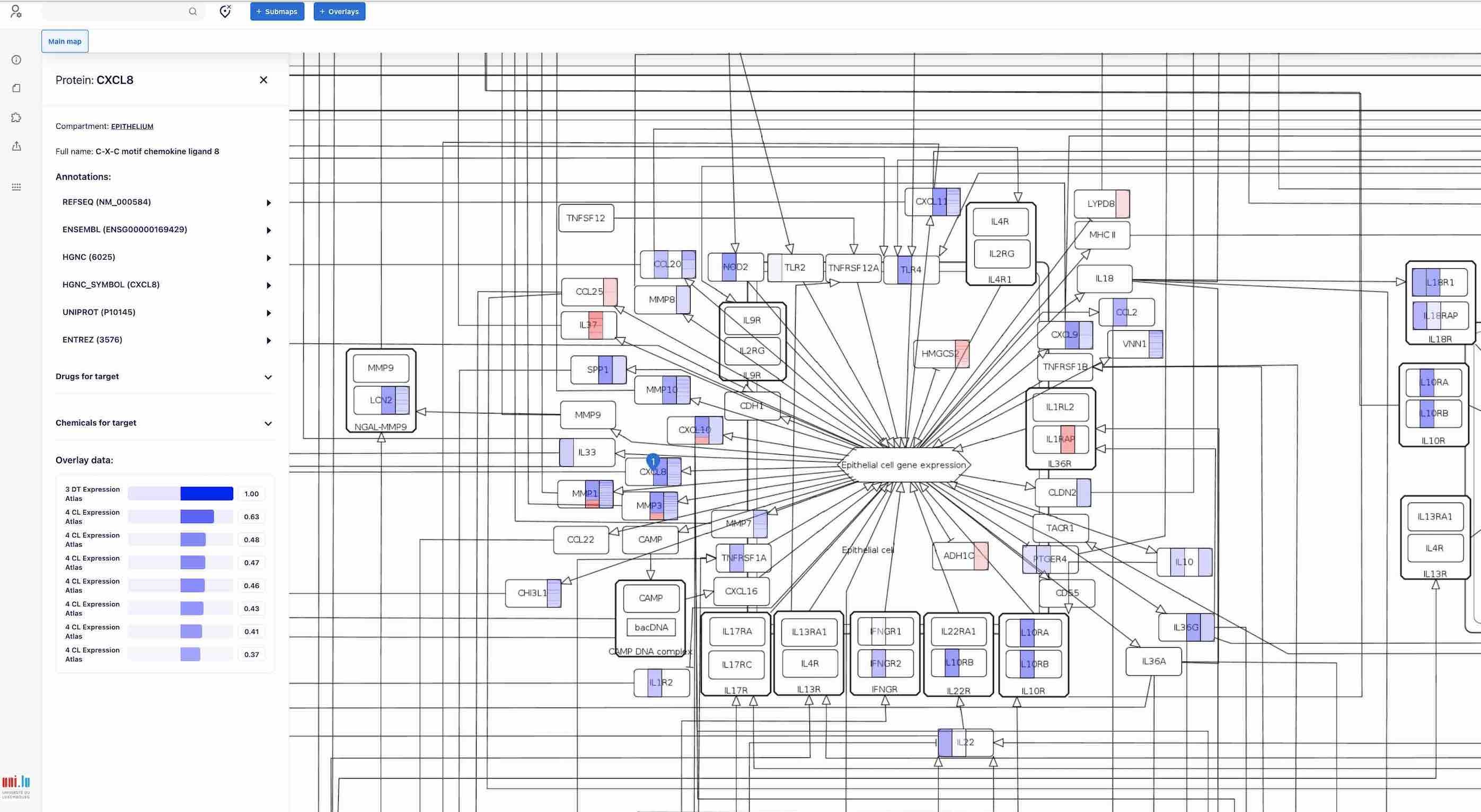
To view details, please click on an element and access the panel on the left with the description.
Supplementary tables
Supplementary Table S5: A complete list of overlapping groups in UC and AD disease maps. The table shows the content of the orverapping groups and provides direct links to them in corresponding disease maps.
Supplementary Table S6: Visualisation of the overlapping pathology mechanisms of UC and AD. Specific connections can be accessed using the URLs provided.